About HyperIDR
Intrinsically disordered regions (IDRs) are pervasive in proteins and play essential roles in diverse biological processes. Accurate identification of these regions is crucial for understanding protein structure and function relationships. Previous studies indicated that short and long IDRs exhibit distinct sequence and functional characteristics. However, existing computational approaches often fail to explicitly model the prevalent scenario in which a single protein harbors multiple coexisting short- and long-disordered regions, thereby limiting predictive accuracy and generalizability. Effectively capturing such heterogeneous disorder semantic within individual proteins remains a significant challenge.
In this study, inspired by the natural analogy between the genotype (hypernetwork) and phenotype (main network), we propose HyperIDR, a multi-scale semantic hypernetwork for modeling heterogeneous disorder semantics across short and long IDRs within individual proteins. Specifically, HyperIDR constructs a semantic prompt pool from representations learned on short and long disordered segments. For a given protein sequence, a prompt-routing module dynamically selects the top-k prompts to capture sequence-level “genotypic” semantics. These prompts guide a hypernetwork to generate parameters for three scale-specific “phenotypic” networks, namely short-, general-, and long-scale networks. The outputs of these networks are integrated through a gating mechanism and refined by a context-aware encoder to produce final disorder predictions. Extensive experiments on five independent benchmark datasets demonstrate that HyperIDR consistently outperforms or achieves performance comparable to existing predictors while maintaining robust accuracy across proteins with varying proportions of short and long IDRs.
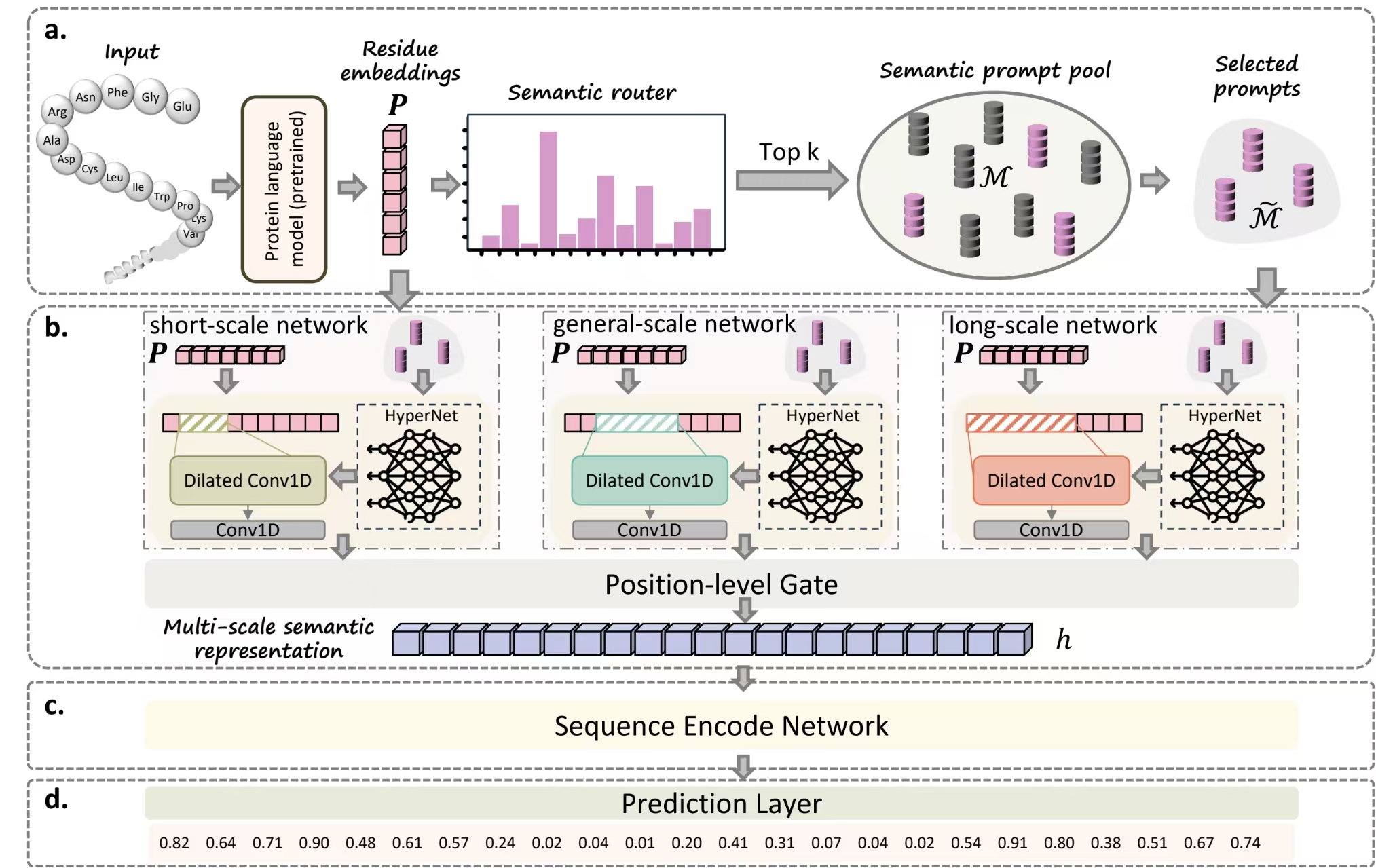
Overview of the HyperIDR framework. (a) semantic prompt routing module dynamically selects top-k semantically relevant prompts from the Semantic Prompt Pool (SPP) and feeds them into a lightweight hypernetwork to generate network parameters; (b) dynamic multi-scale semantic networks module include short-, long-, and general-scale networks that capture multi-scale semantics patterns and are fused by a gating head; (c) sequence encoder (e.g., BiLSTM) contextualizes sequence representations; and (d) prediction module outputs residue-level probabilities of intrinsic disorder.