1. Model and Analysis Options
Model: KGIPA-Fast | SHAP Analysis: True
2. Prediction Results
2.2 Predicted binding residues on peptide and protein sides
2.2.1 Predicted results on peptide side
| 1 | 26 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| E | E | Q | W | A | R | E | I | G | A | Q | L | R | R | M | A | D | D | L | N | A | Q | Y | E | R | R | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | / | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
2.2.2 Predicted results on protein side
| 1 | 50 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| V | L | P | G | E | V | L | A | I | E | G | I | F | M | A | C | G | L | N | E | P | E | Y | L | Y | H | P | L | L | S | P | I | K | L | Y | I | T | G | L | M | R | D | K | E | S | L | F | E | A | M | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| L | A | N | V | R | F | H | S | T | T | G | I | N | Q | L | G | L | S | M | L | Q | V | S | G | D | G | N | M | N | W | G | R | A | L | A | I | L | T | F | G | S | F | V | A | Q | K | L | S | N | E | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 0 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| P | H | L | R | D | F | A | L | A | V | L | P | A | Y | A | Y | E | A | I | G | P | Q | W | F | R | A | R | G | G | W | R | G | L | K | A | Y | C | T | Q | V | L | T | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Legend
Non-Covalent Interactions
Hydrophobic Interactions
Hydrogen Bonds
Water Bridges
Salt Bridges
π Stacks
π Cation Interactions
Halogen Bonds
3. SHAP Analysis based on Non-Covalent Interactions
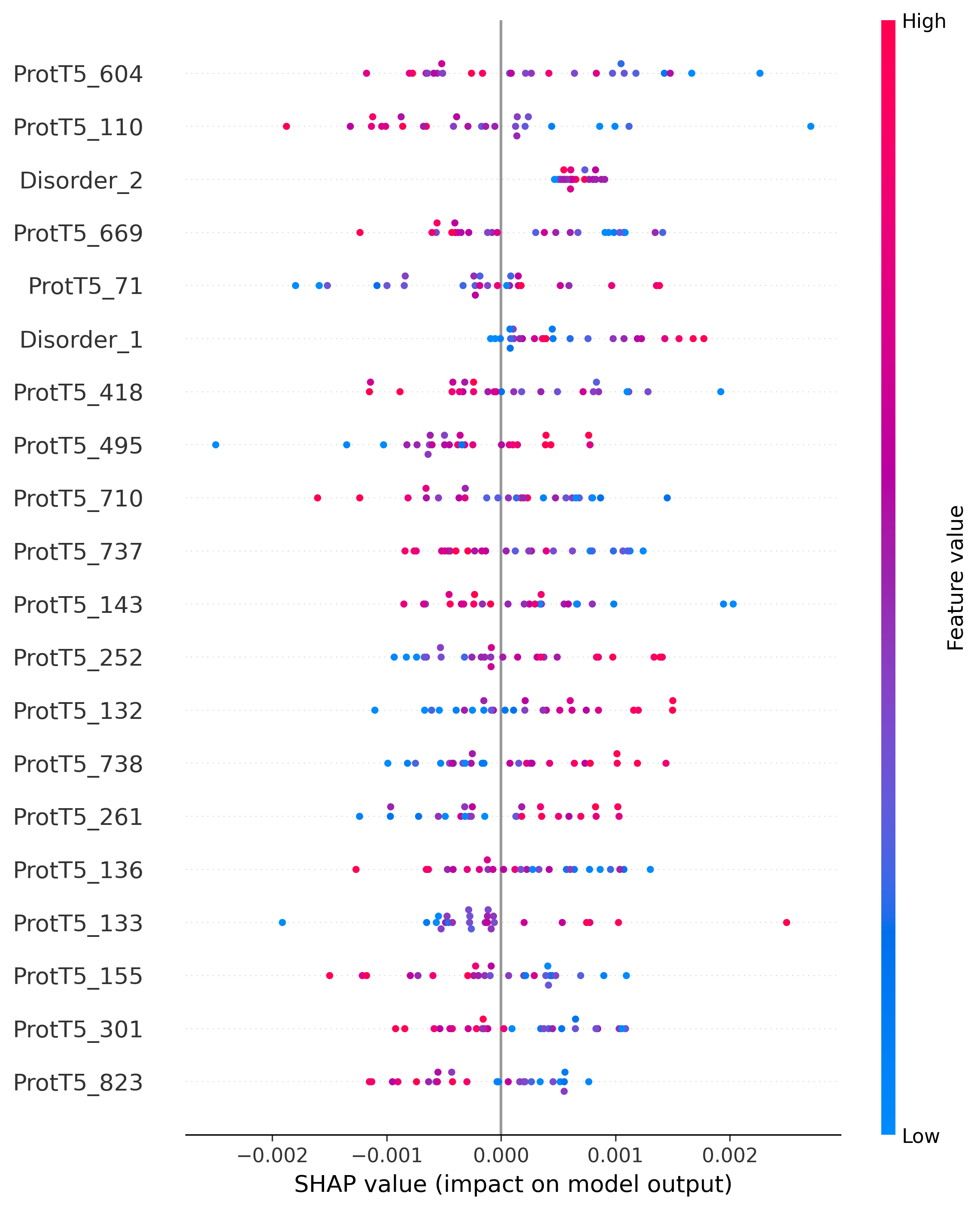
Top 20 Important Peptide-side Features (SHAP Analysis)
Top 20 Important Protein-side Features (SHAP Analysis)
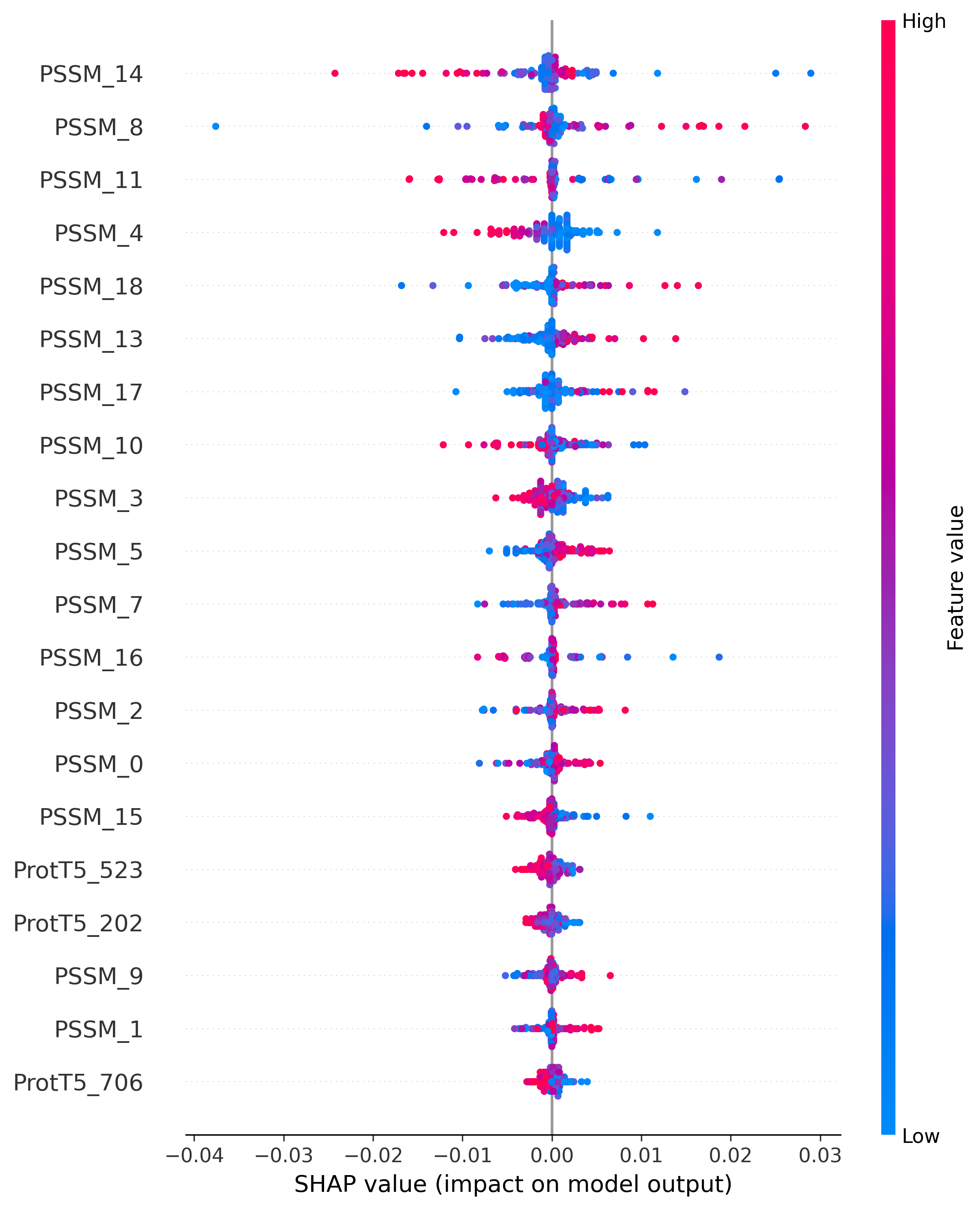
We conducted a feature-level interpretability analysis based on SHAP to evaluate the contribution of each feature type to the model’s prediction.
For different non-covalent interaction types, the SHAP analyses of peptide-side and protein-side features were performed separately,
facilitating researchers to reproduce results similar to Fig. 4B in our paper.
Specifically,
Integer_encoding denotes integer encoding,
Physicochemical refers to physicochemical properties,
Disorder corresponds to disorder tendency,
PSSM indicates evolutionary information,
ProtT5 denotes pretrained embeddings,
and ProtT5 represents pretrained embeddings from large language models.
The numeric suffix following each feature name corresponds to the feature dimension.
Our analysis indicates that the model places greater weight on pretrained embeddings and disorder propensity on the peptide side,
whereas on the protein side the main contributors are PSSM-derived evolutionary information and pretrained embeddings.
Download the complete SHAP analysis results, together with the peptide and protein feature matrices:
Download4. Download Features
4.1 Peptide feature download
| Feature Name | Description | Download |
|---|---|---|
| Disorder Trend (Long) | Long-range disorder tendency | Download |
| Disorder Trend (Short) | Short-range disorder tendency | Download |
| Disorder Trend (Glob) | Global disorder tendency | Download |
| Pretrained Embeddings (ProtT5) | Protein language model embeddings | Download |
4.2 Protein feature download
| Feature Name | Description | Download |
|---|---|---|
| Disorder Trend (Long) | Long-range disorder tendency | Download |
| Disorder Trend (Short) | Short-range disorder tendency | Download |
| Disorder Trend (Glob) | Global disorder tendency | Download |
| Pretrained Embeddings (ProtT5) | Protein language model embeddings | Download |
| Evolutionary Trend (PSSM) | Position-specific scoring matrix | Download |
Note:
- Disorder trends are predicted by IUPred2A (long, short, and glob).
- Pretrained embeddings are from ProtT5.
- PSSM features are generated using PSI-BLAST based on nrdb90, with missing positions filled using BLOSUM substitution scores.
- Disorder trends are predicted by IUPred2A (long, short, and glob).
- Pretrained embeddings are from ProtT5.
- PSSM features are generated using PSI-BLAST based on nrdb90, with missing positions filled using BLOSUM substitution scores.
Please download or bookmark this page (Ctrl + D) to access your results later.
Analysis results will be retained for 7 days only.