Introduction
Motivation: Precision oncology requires resolving intratumoral heterogeneity (ITH) to eliminate minimal residual disease (MRD), a primary driver of therapeutic failure. Although single-cell RNA sequencing (scRNA-seq) enables detection of rare resistant subpopulations, drug-response prediction is hindered by extreme sparsity, noise, and class imbalance. Existing deep learning models are often "black boxes" that lack pharmacological constraints and do not explicitly align drug mechanisms with cellular pathways, limiting interpretability and actionable insight into resistance.
Results: We present scRADAR (Response Analysis via Drug-Aware Routing), a mechanism-guided prototype network that couples accurate prediction with biological transparency. scRADAR builds a robust dual-view cellular representation by integrating metabolic and signaling pathway activities, and uses feature-wise linear modulation (FiLM) to condition cellular embeddings on drug mechanisms. Through sparse prototype routing, scRADAR decomposes predictions into interpretable response archetypes rather than opaque scores, enabling inspection of resistance-associated programs. Across nine independent cohorts, scRADAR achieves state-of-the-art performance and superior generalization, particularly for identifying rare resistant clones under severe class imbalance. Mechanistic analyses highlight context-dependent resistance patterns, including TGF-β–associated EMT in Erlotinib resistance and metabolic reprogramming in BET-inhibitor resistance, providing a high-resolution framework for hypothesis generation in precision oncology.
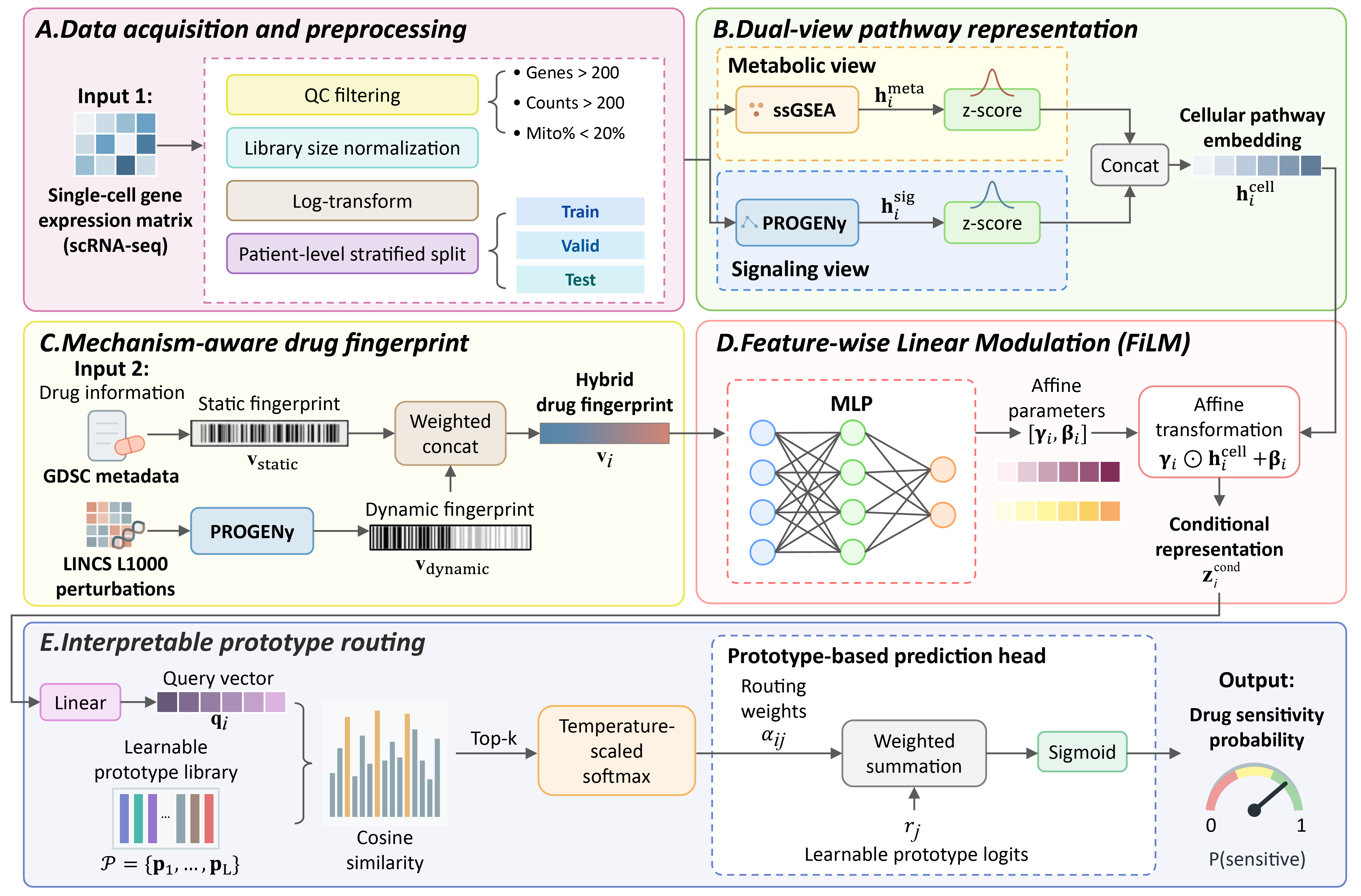
Fig. 1. Overview of the scRADAR framework. (A) scRNA-seq data are QC-filtered, library-size normalized, log-transformed, and split with patient-level stratification. (B) Dual-view pathway encoding concatenates ssGSEA (metabolic) and PROGENy (signaling) scores into a unified cellular embedding. (C) Mechanism-aware drug fingerprints fuse static GDSC descriptors with dynamic LINCS L1000 perturbation signatures. (D) FiLM conditions cellular embeddings via drug-derived affine parameters to generate a mechanism-conditional representation. (E) Prototype routing computes cosine similarities to learnable prototypes, applies sparse top-k selection, and outputs drug sensitivity probability via similarity-weighted aggregation.