The spatial organization of multicellular ecosystems underpins tissue homeostasis and disease progression. Given the high spatial heterogeneity of complex diseases, accurately identifying condition-specific microenvironments linked to clinical phenotypes is a prerequisite for uncovering disease-driving mechanisms and formulating precision medicine strategies. However, current spatial omics computational methods are largely limited to descriptive spatial clustering; they lack the interpretability required to explore pathological associations and struggle to map complex cellular states within microenvironments directly to macroscopic clinical outcomes. Here, we present DREAM (Dual-stream Representation & Explicit Attribution Modeling), a computational framework for interpretable attribution via concept-driven modeling. DREAM leverages context-aware semantic transfer to construct robust niche semantic representations, synergistically encoding intrinsic biological semantics and extrinsic spatial topology through a dual-stream architecture. By incorporating a concept bottleneck mechanism, the framework maintains a balance between clinical predictive accuracy and biological interpretability. Extensive benchmarking across five multi-scale spatial transcriptomic and proteomic datasets demonstrates DREAM’s superior performance in identifying highly reproducible tissue domains. Applied to clinical cohorts, DREAM accurately predicted slice-level disease states and explicitly identified the microenvironmental drivers of colorectal cancer metastasis by pinpointing the spatial convergence of tumor stemness and stromal remodeling. Furthermore, in liver cancer, the framework autonomously localized functional tertiary lymphoid structures (TLS) in a fully unsupervised manner, yielding a compact, highly prognostic spatial biomarker. Ultimately, DREAM demonstrates how concept-driven modeling can highlight potential pathological associations from complex spatial omics data, providing a novel computational perspective for understanding spatial heterogeneity in complex diseases.
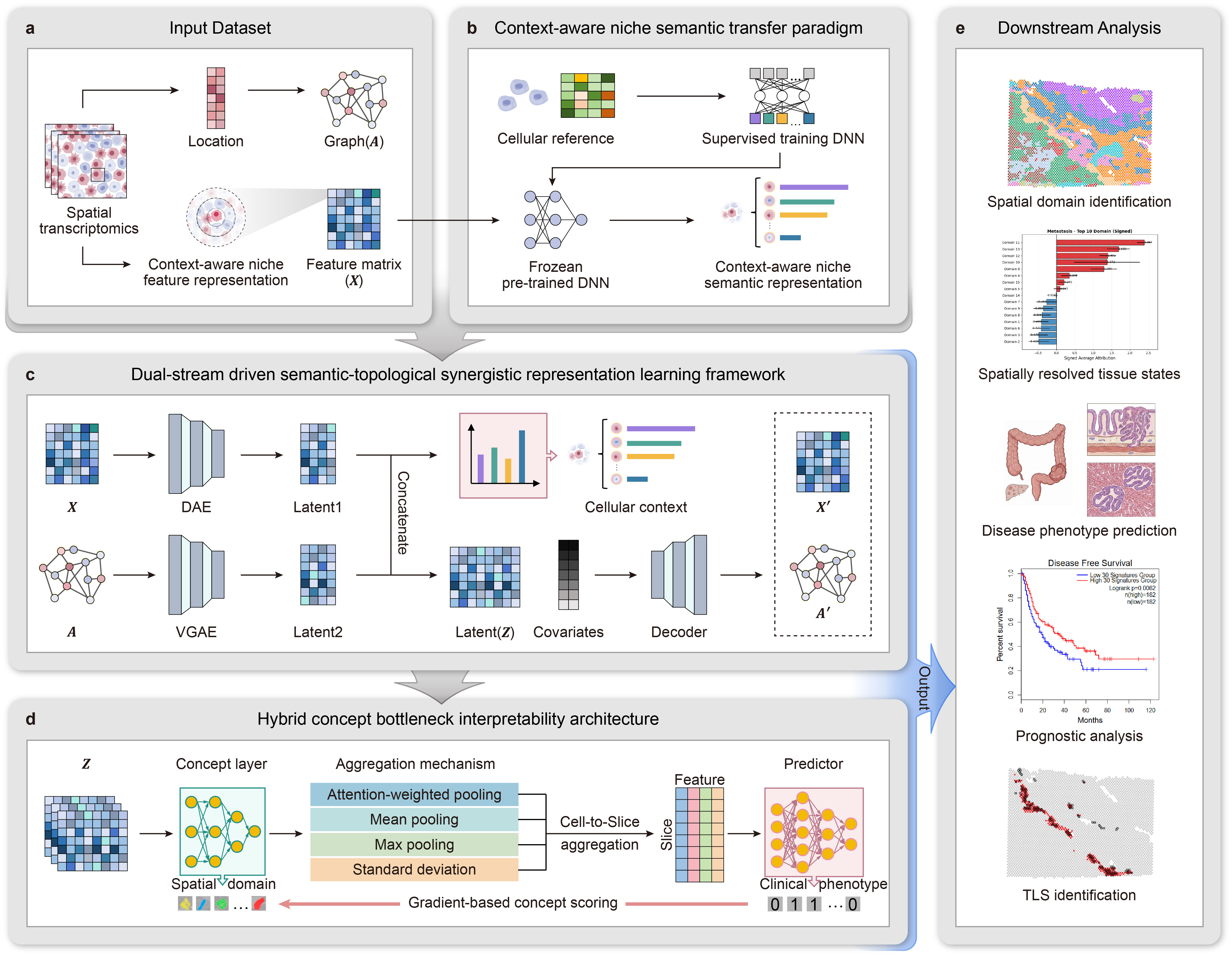
Figure.1 Overall architecture of DREAM. a The workflow begins with the Input dataset module. b The Context-aware niche semantic transfer paradigm (CAST) leverages neighborhood aggregation to transform context-aware niche feature matrices into neighborhood-scale niche semantic representations. c The Dual-stream driven semantic-topological synergistic representation learning framework reconciles feature representation trade-offs. It synergistically encodes biological semantics through a Semantic Stream and spatial topology through a Topology Stream. d The Hybrid concept bottleneck interpretability architecture functions by first mapping fused features to intermediate “concepts” (spatial domains), subsequently constructing a predictor for clinical phenotypes, and allowing for the explicit quantification of domain contributions via gradient back-propagation. e Downstream Analysis utilizes the interpretable model to quantitatively dissect pathological mechanisms, identifying which condition-specific microenvironments drive disease progression and distilling clinically actionable insights.
All analyzed datasets are publicly available and can be accessed via the following links: (1) the mouse spleen CODEX dataset [https://data.mendeley.com/ datasets/zjnpwh8m5b/1]; (2) the human UTUC IMC dataset [https://doi.org/10.5281/zenodo.6376766]; (3) the mouse V1 neocortex STARmap dataset [https://zenodo.org/record/ 7830764#.ZDpObi-1HUI]; (4) the MERFISH Frontal cortex dataset [https://cellxgene.cziscience.com/ collections/31937775-0602-4e52-a799-b6acdd2bac2e]; (5) the human DLPFC 10X Visium dataset [http://spatial.libd.org/spatialLIBD/] [https://www.ncbi.nlm.nih.gov/geo/]; (6) the CRCLM dataset can be accessed via the link [https://drive.google.com/file/d/1QsQIT0iwcWBFzUBcLUPKYuSnSBxfaME/view?usp=drive_link] [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE132465]; (7) the liver datasets [https://ngdc.cncb.ac.cn/gsa-human/browse/HRA000437] [https://db.cngb.org/search/project/CNP0000650].
We provided codes for reproducing the experiments of the paper "Interpretable attribution of clinical phenotypes to condition-specific microenvironment via concept-driven modeling", and comprehensive tutorials for using DREAM. Please check the tutorial website for more details.
Upon the usage the users are requested to use the following citation:
· D. Zhang, R. Qi, and B. Liu, "Interpretable attribution of clinical phenotypes to condition-specific microenvironment via concept-driven modeling,"