 |
DeepDRBP-2L:a new genome annotation predictor for identifying DNA-binding proteins and RNA-binding proteins using Convolutional Neural Network and Long Short-Term Memory |
 |
DeepDRBP-2L:a new genome annotation predictor for identifying DNA-binding proteins and RNA-binding proteins using Convolutional Neural Network and Long Short-Term Memory |
DeepDRBP-2L is constructed by combining CNN and LSTM for identifying DNA-binding protein, RNA-binding protein and DNA/RNA-binding protein, which is compatible with most major browsers, and the parallel speed-up is implemented. It offers an open and interactive web service, which accepts query sequences in FASTA format and returns the search results in a user-friendly manner. For the convenience of the experimental scientists, a step-by-step guide on how to use the DeepDRBP-2L web server is given below.
Visit the web server by clicking the link at http://bliulab.net/DeepDRBP-2L/server and you will see the page as shown in Fig. 1. The Microsoft Edge and Google Chrome browsers are recommended.
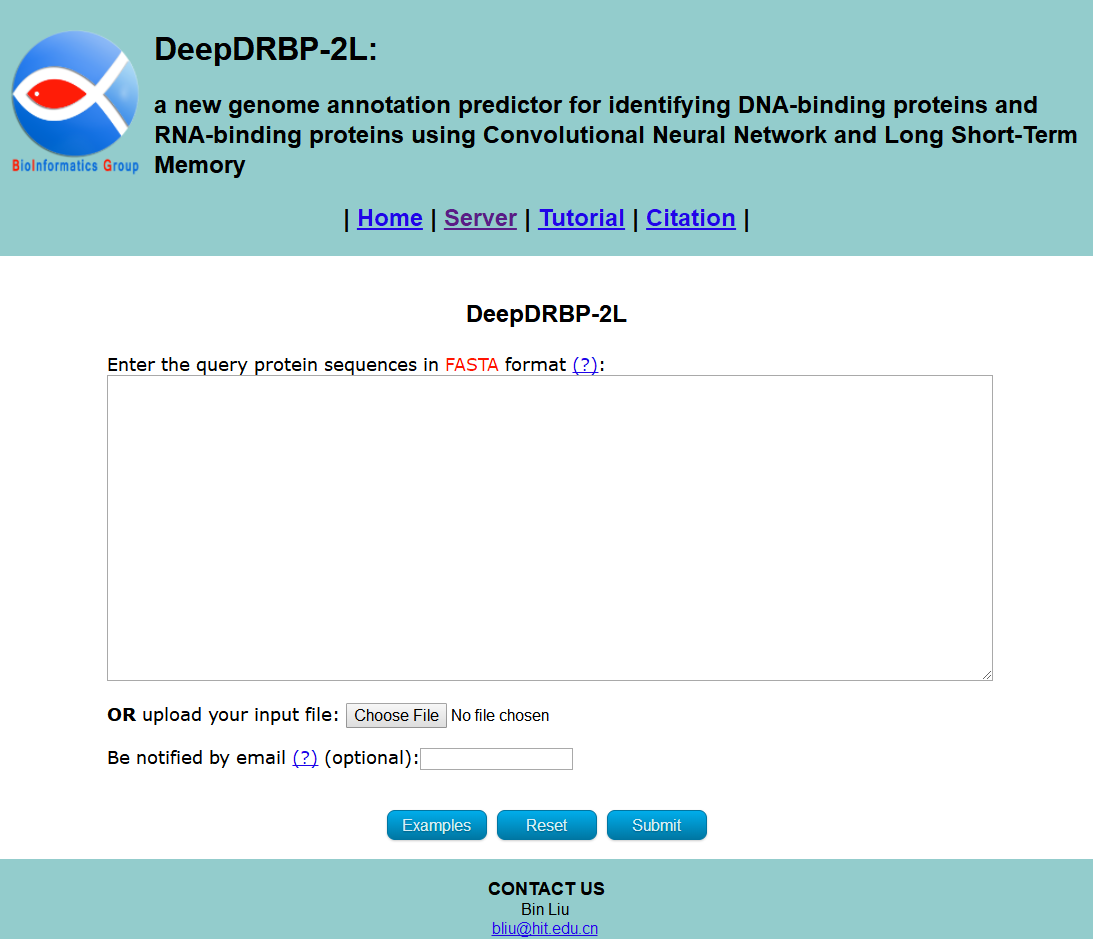
STEP 1: Input the query protein sequences
You can directly enter/paste the query protein sequences into the input box, or upload them via file by clicking the Choose File button. All the input sequences should be in the FASTA format. A sequence in FASTA format consists of a single line beginning with the symbol ">" and multiple lines of amino acids data. You can click the Examples button to automatically input the built-in sequence examples, as show in Fig. 2, and click the Reset button to empty all input sequences. The Email address is optional, please input your valid Email address so that we can send the results to you.
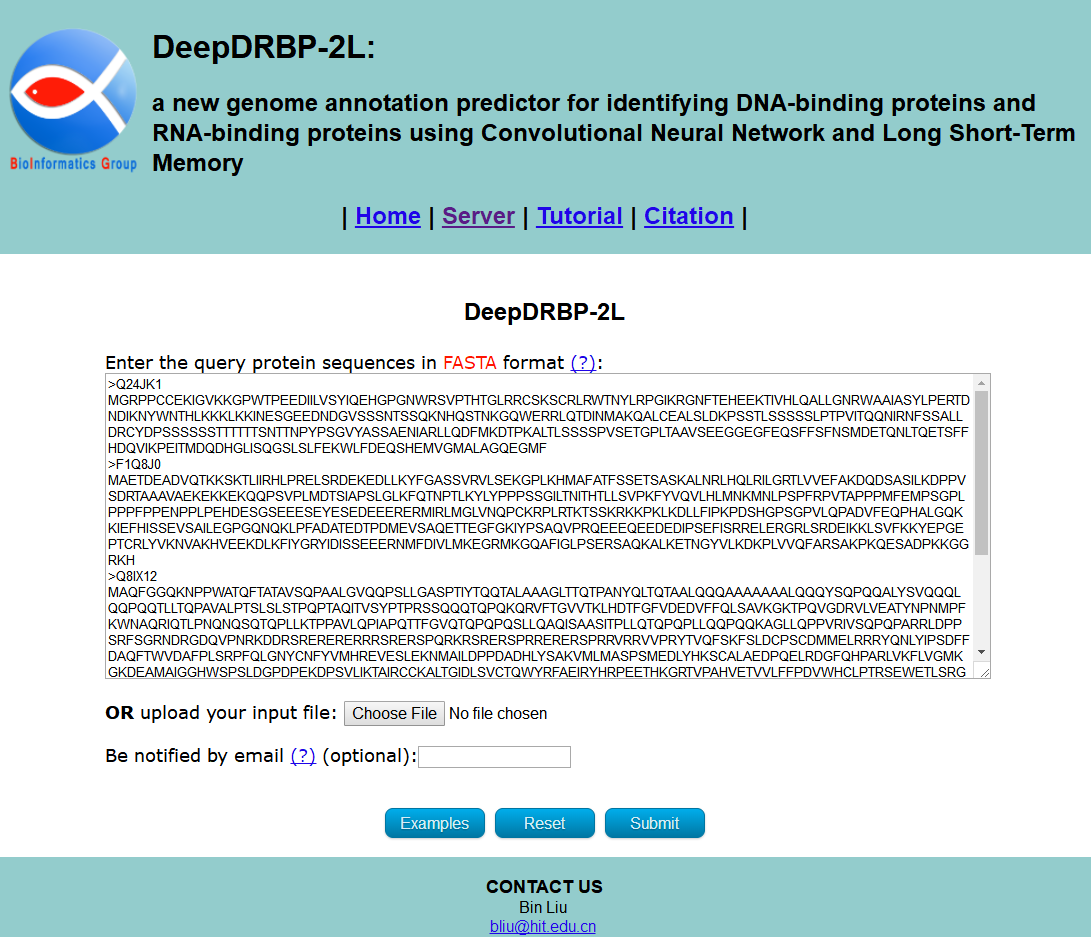
STEP 2: Submit your queries
If you enter query protein sequences, and then click the Submit button, you will see a processing page as shown in Fig. 3. Your job is being processed. The results will be shown on your screen when it is finished. You can also close this browser window, and reload the results by using the link.
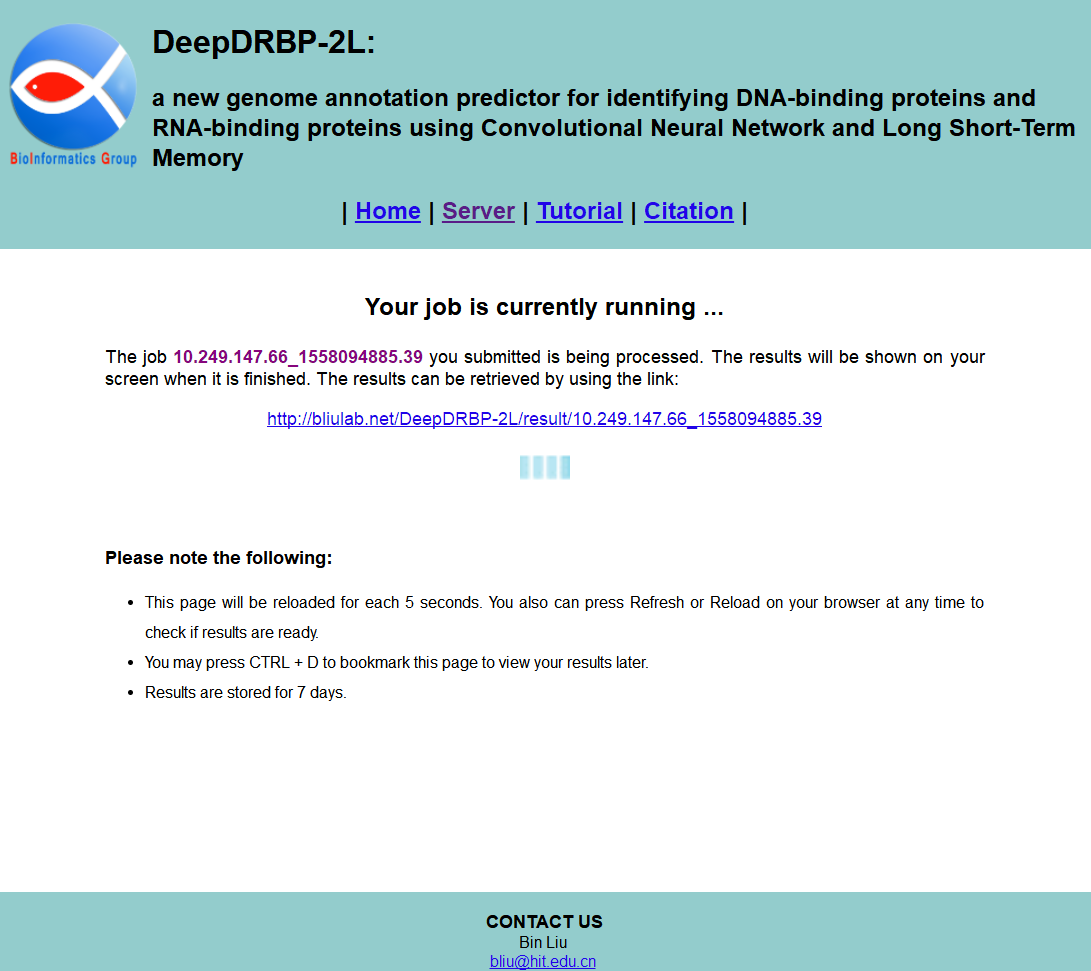
STEP 3: View results
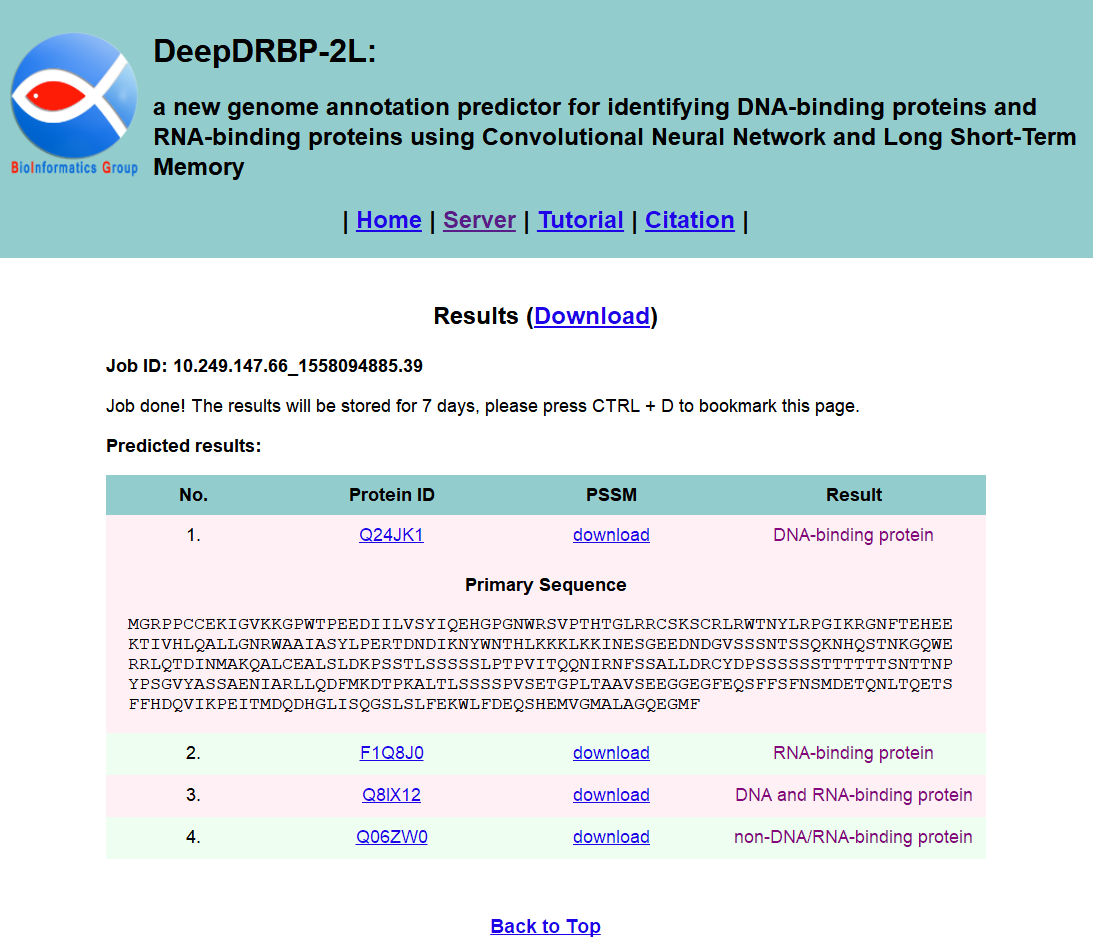
The results will be shown on your screen when it is finished, as shown in Fig. 4. The results will be stored for 7 days, and you can click the Download link to download it. You can find the protein primary sequence by clicking the Protein ID button in Visualization column of the table. Corresponding Position Specific Scoring Matrix (PSSM) can be download by clicking the corresponding download buttons in Visualization column of the table.
CONTACT US
Bin Liu
bliu@bliulab.net
School of Computer Science and Technology, Beijing Institute of Technology, China.
网站备案号: 粤ICP备19041859号-1
Copyright@ By Liu Lab, Beijing Institute of Technology.