 |
PreRBP-TL:prediction of species-specific RNA-binding proteins based on transfer learning |
 |
PreRBP-TL:prediction of species-specific RNA-binding proteins based on transfer learning |
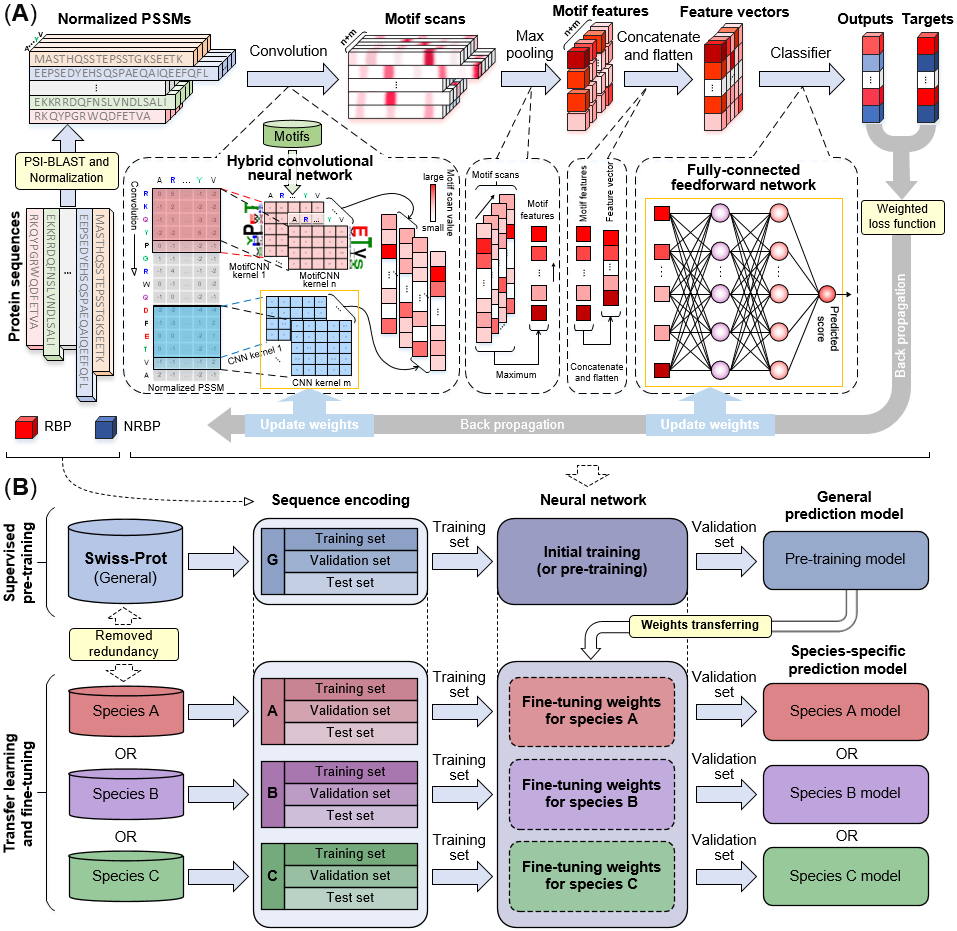
PreRBP-TL is a new sequence-based computational predictor for identifying species-specific RNA-binding proteins (RBPs). In which hybrid convolutional neural networks (CNNs) and transfer learning were employed. The weights of prediction model were initialized by pre-training with the large general RBP dataset, and then fine-tuned with the small species-specific RPB dataset by using transfer learning. The experimental results show that the PreRBP-TL achieves obviously higher performance for identifying the species-specific RBPs from six different organisms, outperforming eight state-of-the-art computational methods, which is an important complement to existing methods. The network architecture and framework of PreRBP-TL are shown in Fig. 1. For more information please refer to our paper.
The structural motifs used in this study: motifs.txt
Pfam RNA-binding domains used in this study: RBDs.txt
All the datasets used in this study: Datasets.zip
We acknowledge with thanks the following softwares used as a part of this server:
(1) PSI-BLAST - Generation of the position specific scoring matrices (PSSMs);
(2) NRDB90 - The non-redundant database for usage of PSI-BLAST to generate PSSMs;
(3) Biopython - Protein data processing;
(4) Keras - Construction of the prediction model;
(5) Tensorflow - Backend of keras for calculation.
Upon the usage of this server the users are requested to use the following citation:
Zhang, J., Yan, K., Chen, Q. and Liu, B. PreRBP-TL: prediction of species-specific RNA-binding proteins based on transfer learning. Bioinformatics, 2022, 38(8): 2135-2143.
CONTACT US
Bin Liu
bliu@bliulab.net
School of Computer Science and Technology, Harbin Institute of Technology, Shenzhen, Guangdong 518055, China.
Copyright © by Liu Lab, Harbin Institute of Technology
网站备案号:粤ICP备19041859号-1