 |
ProtDec-LTR 2.0An improved method for protein remote homology detection by combining pseudo protein and supervised learning to rank |
 |
ProtDec-LTR 2.0An improved method for protein remote homology detection by combining pseudo protein and supervised learning to rank |
ProtDec-LTR2.0 web server is constructed based on SCOPe v2.06, which is compatible with most major browsers, and the parallel speed-up is implemented. It offers an open and interactive web service, which accepts query sequences in FASTA format and returns the search results in a user-friendly manner. For the convenience of the experimental scientists, a step-by-step guide on how to use the ProtDec-LTR 2.0 web server is given below.
Visit the web server by clicking the link at http://bliulab.net/ProtDec-LTR2.0/server and you will see the page as shown in Fig. 1. The Microsoft Edge and Google Chrome browsers are recommended.
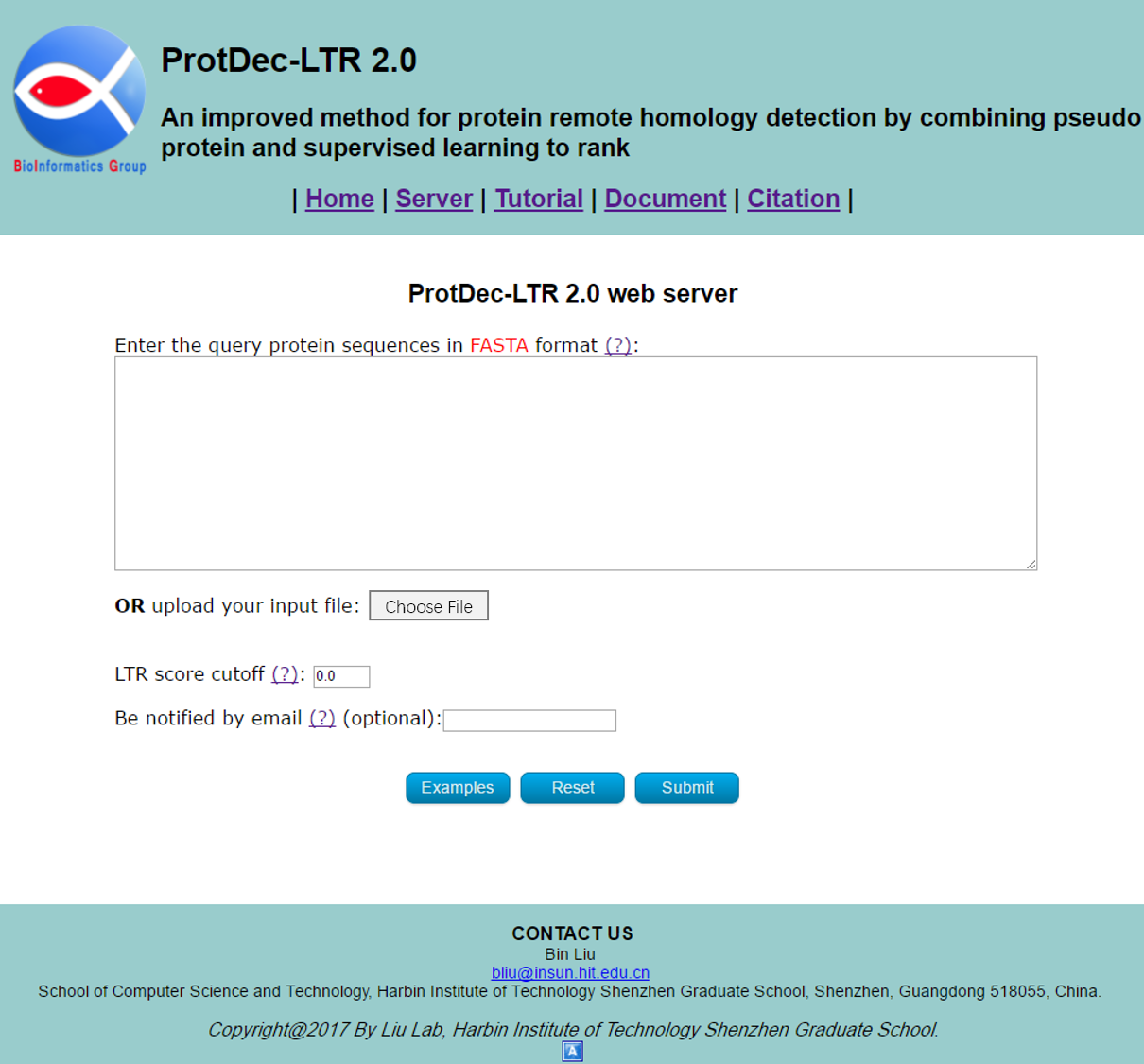
STEP 1: Input the query protein sequences
You can directly enter/paste the query protein sequences into the input box, or upload them via file by clicking the Choose File button. All the input sequences should be in the FASTA format. A sequence in FASTA format consists of a single line beginning with the symbol ">" and multiple lines of amino acids data. You can click the Examples button to automatically input the built-in sequence examples and the default parameter as shown in Fig 2, and click the Reset button to empty all input sequences and parameters.
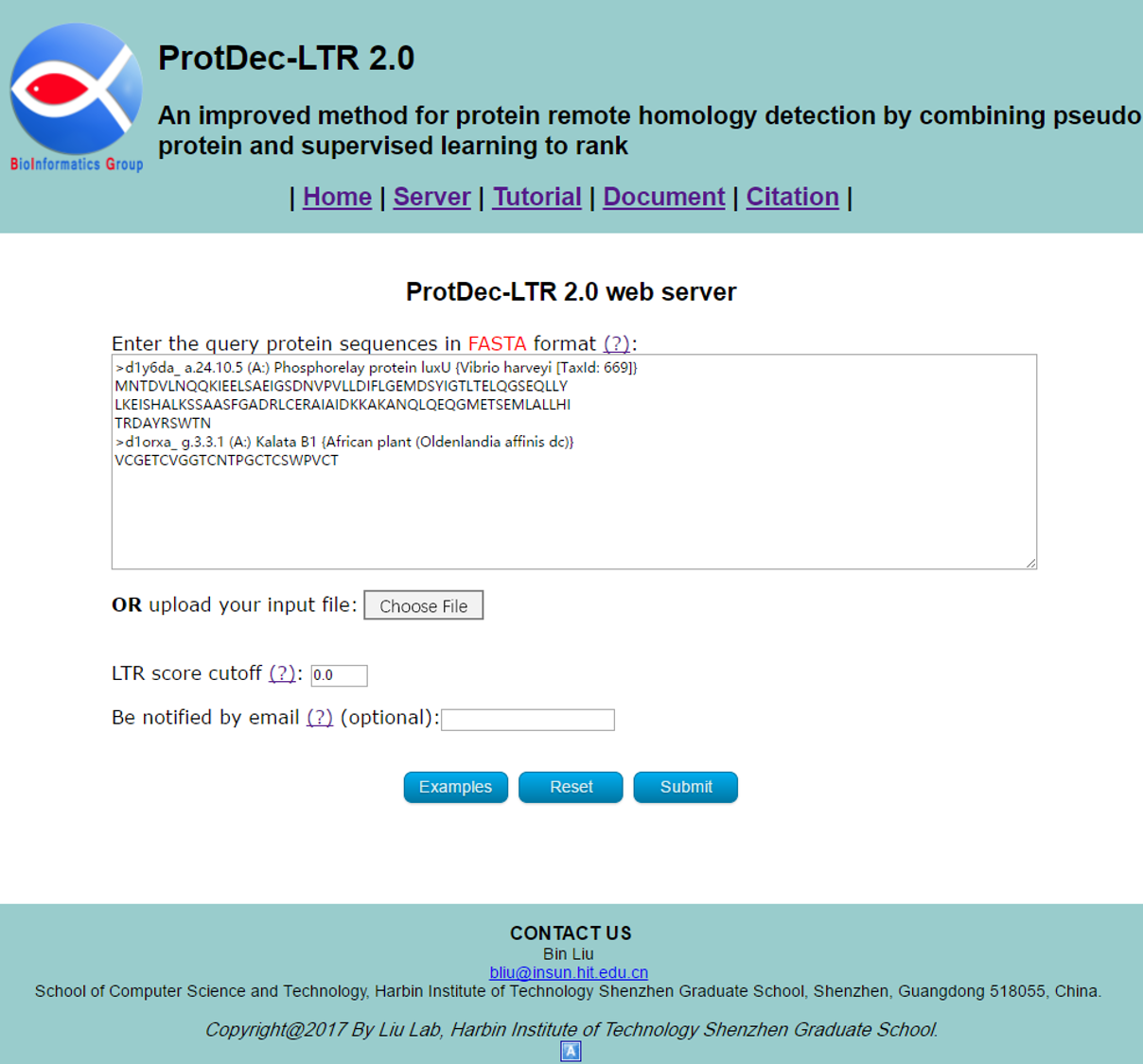
STEP2: Set the parameters
The LTR score cutoff is necessary, which is a threshold for results output. Its default value is 0.0, which is optimized for most users. Besides, the Email address is optional, please input your valid Email address so that we can send the results to you.
STEP 3: Submit your queries
If you enter query protein sequences, and then click the Submit button, you will see a processing page as shown in Fig. 3. Your job is being processed. The results will be shown on your screen when it is finished. You can also close this browser window, and reload the results by using the link.
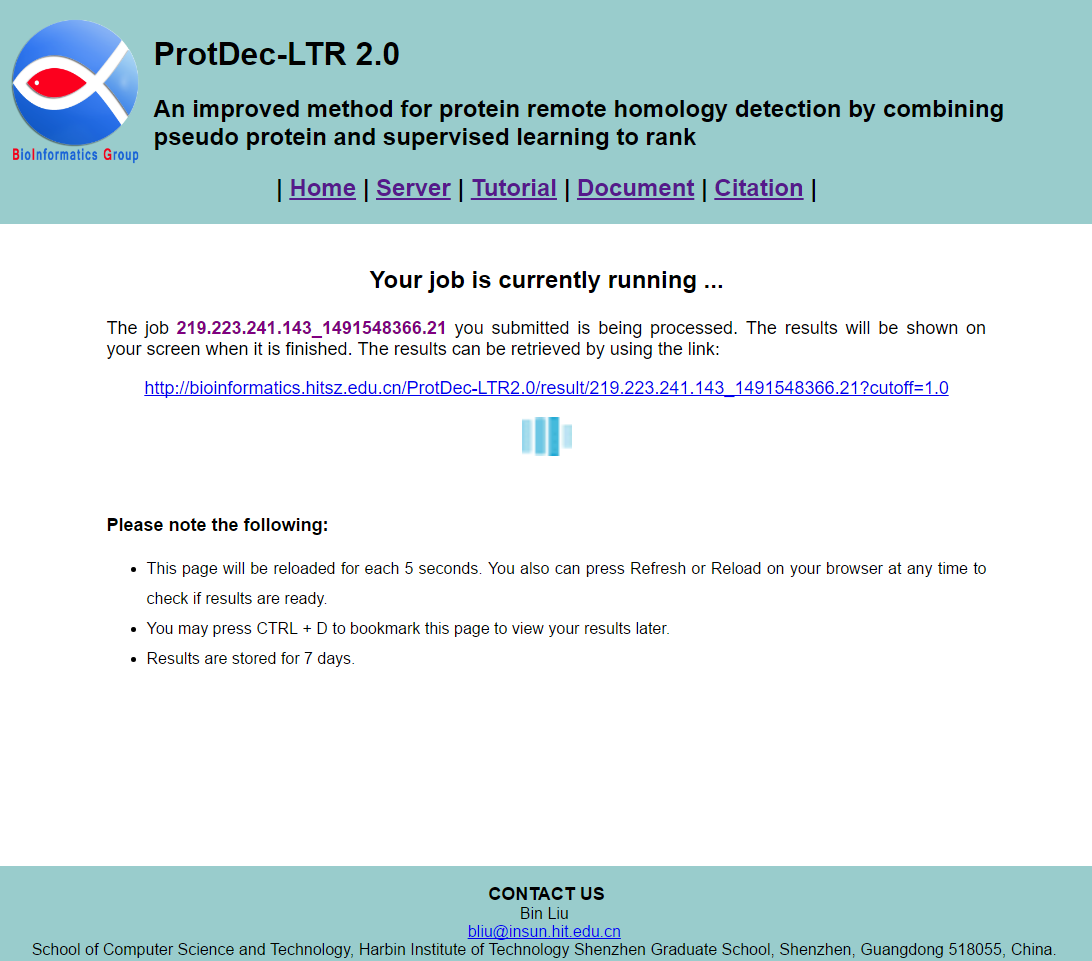
STEP 4: View results
The results will be shown on your screen when it is finished, as shown in Fig. 4. The results will be stored for 7 days, and you can click the Download link to download it. You can click the query IDs to view their corresponding results. For each query, their homologous proteins are listed in a table. Some useful sequence and structure information are provided, including SCOP ID, family classification, sequence length. You can find the protein primary sequence and 3D structure by click the 3D Structure button in Visualization column of the table. The more detailed structure summary and biology knowledge on this protein can be found by clicking the RCSB PDB and SBKB.org button at bottom of this cell.
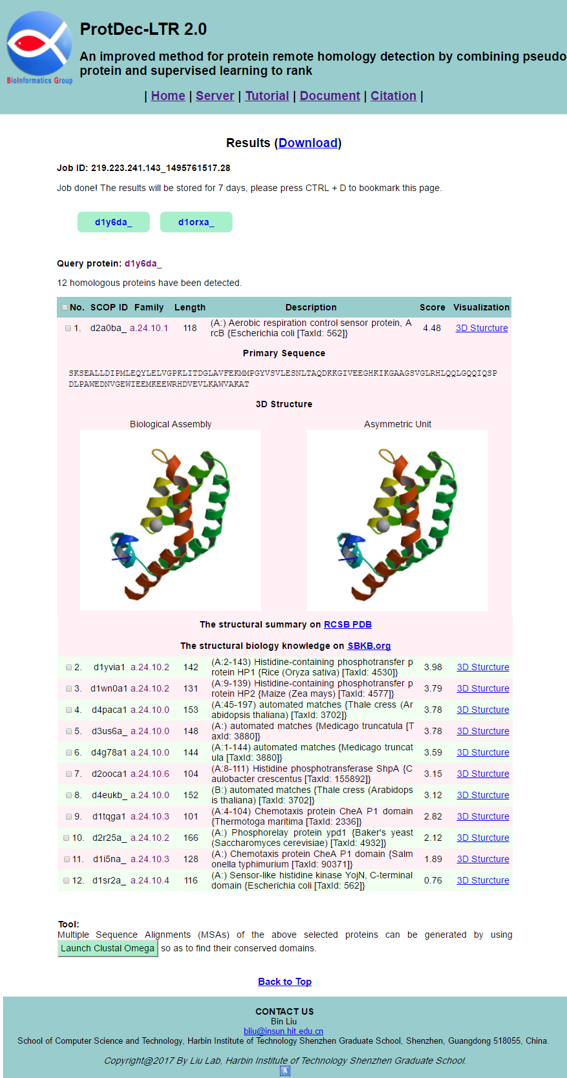
STEP 5: Generate multiple sequence alignment
At the bottom of result page, we provide a tool for generating multiple sequences alignment. First, you should select some sequences by ticking in checkboxes, or select all proteins by ticking the checkbox in the table header. Then, click the Launch Clustal Omega button to submit your selected sequences. You will see the page as shown in Fig. 5. And according this results, you can easily find their conserved domains for further study.
